Find the Orthogonal Trajectories of the Family of Curves. (Use C for Any Needed Constant.) Y2 = 5kx3
3.vi. scikit-learn: auto learning in Python¶
Authors: Gael Varoquaux

Capacity contents
- Introduction: problem settings
- Bones principles of machine learning with scikit-learn
- Supervised Learning: Classification of Handwritten Digits
- Supervised Learning: Regression of Housing Data
- Measuring prediction operation
- Unsupervised Learning: Dimensionality Reduction and Visualization
- The eigenfaces instance: chaining PCA and SVMs
- The eigenfaces instance: chaining PCA and SVMs
- Parameter selection, Validation, and Testing
- Examples for the scikit-learn chapter
3.6.1. Introduction: problem settings¶
3.6.one.1. What is auto learning?¶
Tip
Machine Learning is well-nigh edifice programs with tunable parameters that are adapted automatically and then as to meliorate their behavior past adapting to previously seen data.
Machine Learning can be considered a subfield of Artificial Intelligence since those algorithms can be seen as building blocks to make computers larn to bear more intelligently by somehow generalizing rather that just storing and retrieving data items like a database system would do.
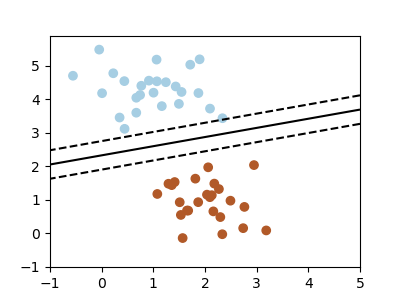
A classification problem
We'll take a look at two very simple auto learning tasks here. The starting time is a classification task: the figure shows a collection of two-dimensional information, colored co-ordinate to 2 unlike class labels. A classification algorithm may exist used to draw a dividing boundary between the two clusters of points:
By drawing this separating line, we have learned a model which can generalize to new data: if you were to drop some other bespeak onto the aeroplane which is unlabeled, this algorithm could at present predict whether it's a blue or a red point.
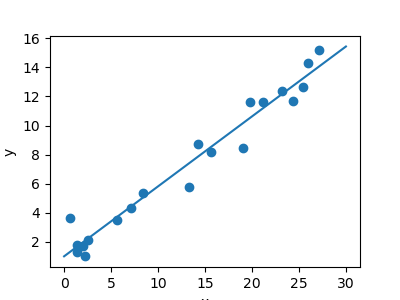
A regression problem
The next simple task we'll look at is a regression task: a elementary best-fit line to a set of data.
Once again, this is an example of plumbing fixtures a model to information, but our focus hither is that the model can make generalizations about new data. The model has been learned from the training data, and can be used to predict the event of test data: here, we might be given an x-value, and the model would allow the states to predict the y value.
three.6.1.2. Data in scikit-larn¶
The data matrix¶
Machine learning algorithms implemented in scikit-learn await data to exist stored in a ii-dimensional array or matrix. The arrays can be either numpy arrays, or in some cases scipy.sparse matrices. The size of the assortment is expected to exist [n_samples, n_features]
- n_samples: The number of samples: each sample is an detail to procedure (eastward.k. classify). A sample tin can be a document, a picture, a sound, a video, an astronomical object, a row in database or CSV file, or whatever you tin can describe with a stock-still set of quantitative traits.
- n_features: The number of features or distinct traits that tin be used to depict each item in a quantitative manner. Features are generally real-valued, but may be boolean or discrete-valued in some cases.
Tip
The number of features must be fixed in advance. However information technology can be very high dimensional (eastward.g. millions of features) with most of them existence zeros for a given sample. This is a example where scipy.sparse matrices can be useful, in that they are much more retentivity-efficient than numpy arrays.
A Unproblematic Case: the Iris Dataset¶
The application problem¶
Equally an example of a simple dataset, let u.s. a await at the iris information stored by scikit-learn. Suppose nosotros desire to recognize species of irises. The data consists of measurements of iii different species of irises:
 |  |  |
|---|---|---|
| Setosa Iris | Versicolor Iris | Virginica Iris |
Quick Question:
If we want to design an algorithm to recognize iris species, what might the data exist?
Remember: we demand a 2D array of size
[n_samples x n_features].
- What would the
n_samplesrefer to?- What might the
n_featuresrefer to?
Remember that there must be a fixed number of features for each sample, and feature number i must be a similar kind of quantity for each sample.
Loading the Iris Data with Scikit-learn¶
Scikit-learn has a very straightforward prepare of data on these iris species. The data consist of the following:
- Features in the Iris dataset:
- sepal length (cm)
- sepal width (cm)
- petal length (cm)
- petal width (cm)
- Target classes to predict:
- Setosa
- Versicolour
- Virginica
scikit-learn embeds a copy of the iris CSV file along with a part to load it into numpy arrays:
>>> from sklearn.datasets import load_iris >>> iris = load_iris () Annotation
Import sklearn Note that scikit-learn is imported as sklearn
The features of each sample flower are stored in the data attribute of the dataset:
>>> print ( iris . data . shape ) (150, 4) >>> n_samples , n_features = iris . data . shape >>> print ( n_samples ) 150 >>> print ( n_features ) 4 >>> print ( iris . data [ 0 ]) [five.1 three.5 ane.4 0.2] The information about the grade of each sample is stored in the target attribute of the dataset:
>>> impress ( iris . target . shape ) (150,) >>> print ( iris . target ) [0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 one ane 1 1 one ane 1 1 ane 1 1 i one one 1 1 1 1 ane i 1 1 i one 1 1 1 1 1 1 1 one i 1 ane one 1 1 i ane 1 1 i one 1 1 1 1 1 1 2 2 two two 2 2 2 two two 2 two 2 ii 2 2 two 2 2 2 two 2 two two 2 ii two 2 ii two 2 two 2 2 2 2 ii 2 2 2 2 two 2 2 2 two 2 ii two 2 2] The names of the classes are stored in the last attribute, namely target_names :
>>> print ( iris . target_names ) ['setosa' 'versicolor' 'virginica'] This data is four-dimensional, only we can visualize 2 of the dimensions at a time using a besprinkle plot:
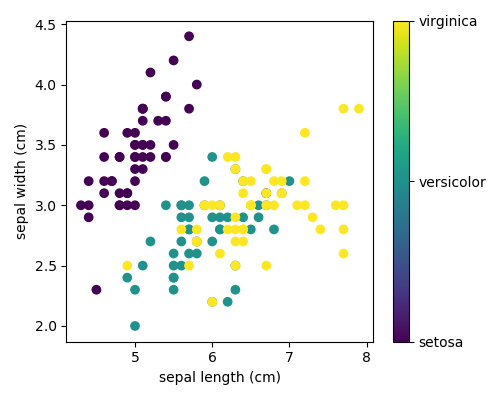
Excercise:
Tin can you choose 2 features to find a plot where it is easier to seperate the different classes of irises?
Hint: click on the effigy above to see the code that generates it, and modify this code.
3.6.two. Bones principles of machine learning with scikit-learn¶
iii.half-dozen.2.1. Introducing the scikit-larn estimator object¶
Every algorithm is exposed in scikit-learn via an ''Computer'' object. For instance a linear regression is: sklearn.linear_model.LinearRegression
>>> from sklearn.linear_model import LinearRegression Estimator parameters: All the parameters of an reckoner tin be set when information technology is instantiated:
>>> model = LinearRegression ( normalize = True ) >>> print ( model . normalize ) True >>> print ( model ) LinearRegression(copy_X=True, fit_intercept=True, n_jobs=one, normalize=True) Plumbing equipment on data¶
Permit's create some simple information with numpy:
>>> import numpy as np >>> x = np . assortment ([ 0 , 1 , ii ]) >>> y = np . assortment ([ 0 , 1 , 2 ]) >>> X = 10 [:, np . newaxis ] # The input information for sklearn is 2D: (samples == three x features == ane) >>> Ten assortment([[0], [1], [2]]) >>> model . fit ( X , y ) LinearRegression(copy_X=Truthful, fit_intercept=Truthful, n_jobs=1, normalize=True) Estimated parameters: When data is fitted with an estimator, parameters are estimated from the information at manus. All the estimated parameters are attributes of the computer object ending past an underscore:
>>> model . coef_ array([ane.]) three.half dozen.2.two. Supervised Learning: Classification and regression¶
In Supervised Learning, we have a dataset consisting of both features and labels. The task is to construct an computer which is able to predict the characterization of an object given the fix of features. A relatively simple case is predicting the species of iris given a set of measurements of its flower. This is a relatively simple task. Some more complicated examples are:
- given a multicolor image of an object through a telescope, make up one's mind whether that object is a star, a quasar, or a galaxy.
- given a photograph of a person, identify the person in the photo.
- given a list of movies a person has watched and their personal rating of the picture show, recommend a listing of movies they would like (So-chosen recommender systems: a famous instance is the Netflix Prize).
Tip
What these tasks have in mutual is that there is ane or more unknown quantities associated with the object which needs to be determined from other observed quantities.
Supervised learning is further cleaved down into two categories, nomenclature and regression. In nomenclature, the label is discrete, while in regression, the label is continuous. For example, in astronomy, the task of determining whether an object is a star, a milky way, or a quasar is a nomenclature trouble: the characterization is from three singled-out categories. On the other mitt, nosotros might wish to judge the age of an object based on such observations: this would be a regression problem, because the characterization (historic period) is a continuous quantity.
Classification: K nearest neighbors (kNN) is one of the simplest learning strategies: given a new, unknown observation, look upwardly in your reference database which ones have the closest features and assign the predominant class. Allow's try it out on our iris classification trouble:
from sklearn import neighbors , datasets iris = datasets . load_iris () 10 , y = iris . information , iris . target knn = neighbors . KNeighborsClassifier ( n_neighbors = 1 ) knn . fit ( 10 , y ) # What kind of iris has 3cm 10 5cm sepal and 4cm x 2cm petal? print ( iris . target_names [ knn . predict ([[ three , 5 , 4 , 2 ]])]) 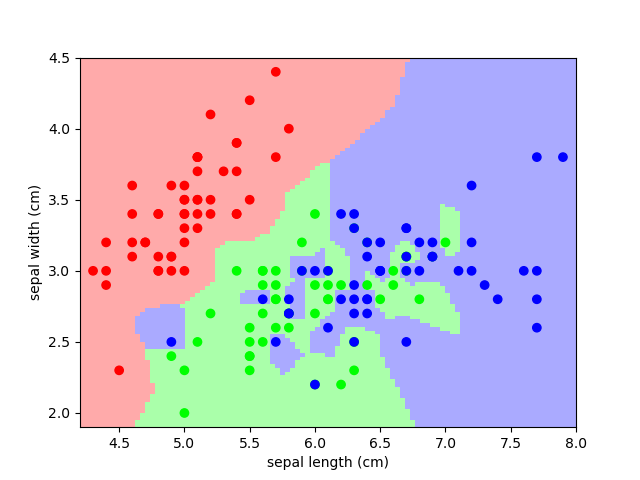
A plot of the sepal infinite and the prediction of the KNN
Regression: The simplest possible regression setting is the linear regression 1:
from sklearn.linear_model import LinearRegression # 10 from 0 to xxx 10 = 30 * np . random . random (( 20 , 1 )) # y = a*10 + b with racket y = 0.five * x + 1.0 + np . random . normal ( size = x . shape ) # create a linear regression model model = LinearRegression () model . fit ( x , y ) # predict y from the data x_new = np . linspace ( 0 , 30 , 100 ) y_new = model . predict ( x_new [:, np . newaxis ]) 
A plot of a simple linear regression.
3.6.ii.3. A recap on Scikit-learn's estimator interface¶
Scikit-larn strives to have a uniform interface across all methods, and nosotros'll see examples of these below. Given a scikit-learn figurer object named model , the post-obit methods are available:
| In all Estimators: | |
|---|---|
| |
| In supervised estimators: | |
| |
| In unsupervised estimators: | |
| |
3.vi.2.4. Regularization: what it is and why information technology is necessary¶
Prefering simpler models¶
Railroad train errors Suppose yous are using a 1-nearest neighbor estimator. How many errors do you wait on your train gear up?
- Train set error is not a good measurement of prediction performance. You demand to exit out a examination set.
- In full general, we should accept errors on the railroad train prepare.
An example of regularization The core thought behind regularization is that we are going to prefer models that are simpler, for a certain definition of ''simpler'', even if they lead to more than errors on the railroad train set.
Every bit an example, let's generate with a 9th order polynomial, with noise:
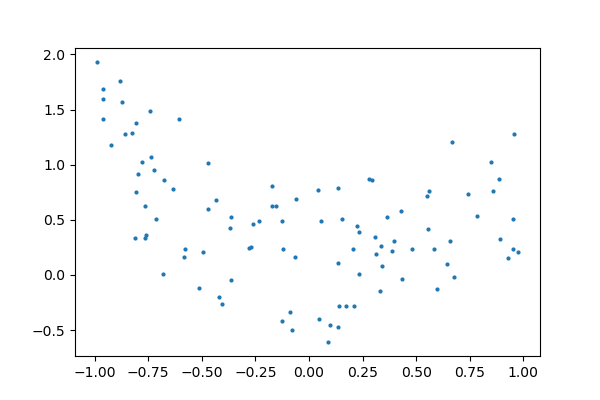
And at present, let's fit a 4th gild and a ninth order polynomial to the data.
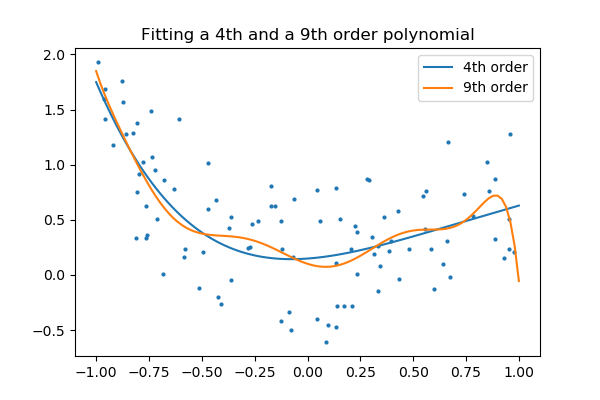
With your naked eyes, which model do you prefer, the 4th order ane, or the 9th gild ane?
Let's look at the basis truth:
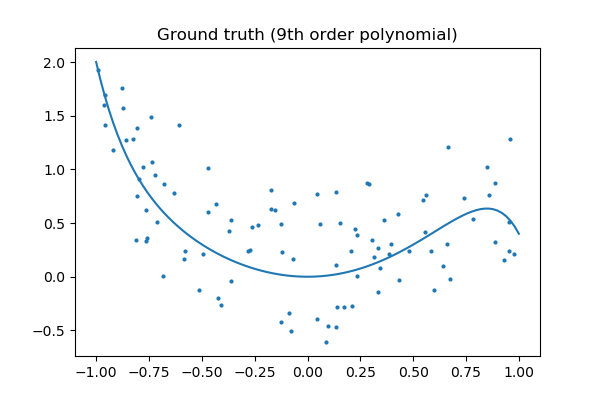
Tip
Regularization is ubiquitous in machine learning. Most scikit-learn estimators have a parameter to tune the amount of regularization. For case, with thou-NN, information technology is 'k', the number of nearest neighbors used to make the conclusion. one thousand=1 amounts to no regularization: 0 error on the training set, whereas large thou will push toward smoother decision boundaries in the feature space.
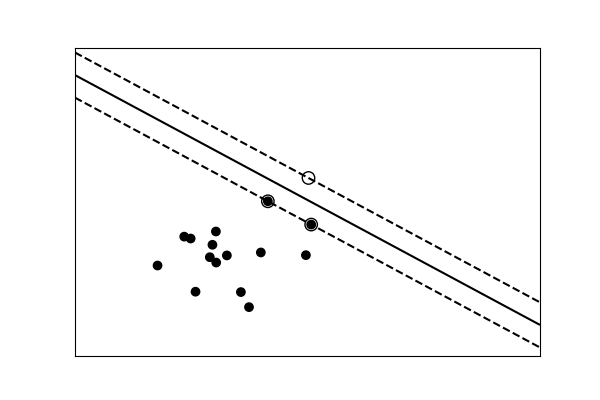
Simple versus complex models for nomenclature¶
 | 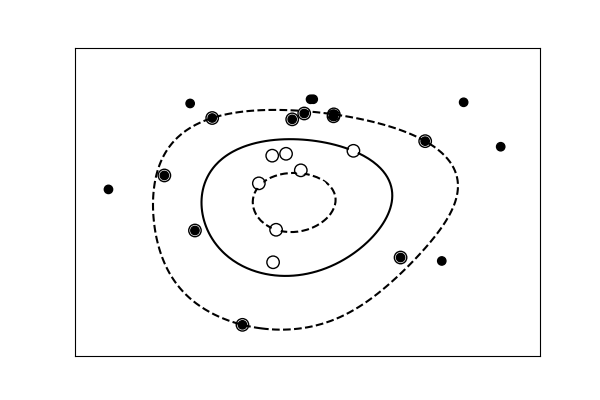 |
|---|---|
| A linear separation | A non-linear separation |
Tip
For classification models, the decision boundary, that separates the class expresses the complexity of the model. For example, a linear model, that makes a decision based on a linear combination of features, is more than circuitous than a non-linear one.
3.half dozen.3. Supervised Learning: Classification of Handwritten Digits¶
3.6.3.1. The nature of the data¶
In this section we'll employ scikit-larn to the classification of handwritten digits. This will become a chip beyond the iris classification nosotros saw before: we'll discuss some of the metrics which tin be used in evaluating the effectiveness of a classification model.
>>> from sklearn.datasets import load_digits >>> digits = load_digits ()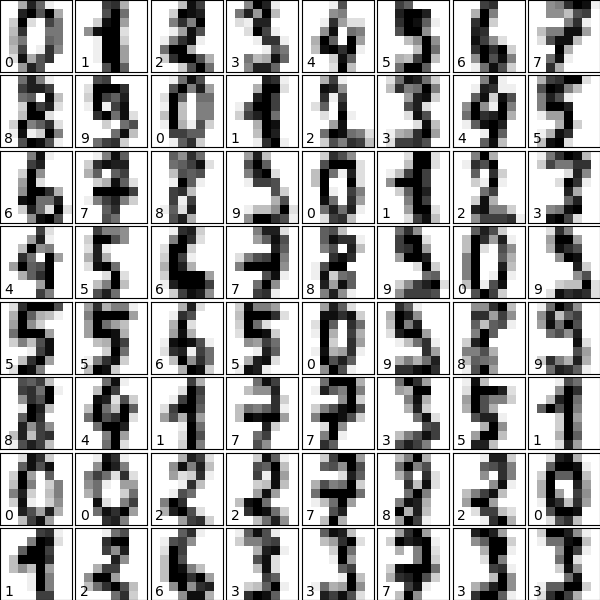 
Let us visualize the data and remind us what nosotros're looking at (click on the figure for the full code):
# plot the digits: each epitome is 8x8 pixels for i in range ( 64 ): ax = fig . add_subplot ( 8 , viii , i + 1 , xticks = [], yticks = []) ax . imshow ( digits . images [ i ], cmap = plt . cm . binary , interpolation = 'nearest' ) iii.6.iii.ii. Visualizing the Data on its primary components¶
A skilful first-step for many problems is to visualize the data using a Dimensionality Reduction technique. We'll start with the most straightforward one, Principal Component Analysis (PCA).
PCA seeks orthogonal linear combinations of the features which show the greatest variance, and as such, tin help requite you a proficient thought of the structure of the data set.
>>> from sklearn.decomposition import PCA >>> pca = PCA ( n_components = ii ) >>> proj = pca . fit_transform ( digits . data ) >>> plt . besprinkle ( proj [:, 0 ], proj [:, ane ], c = digits . target ) <matplotlib.collections.PathCollection object at ...> >>> plt . colorbar () <matplotlib.colorbar.Colorbar object at ...> 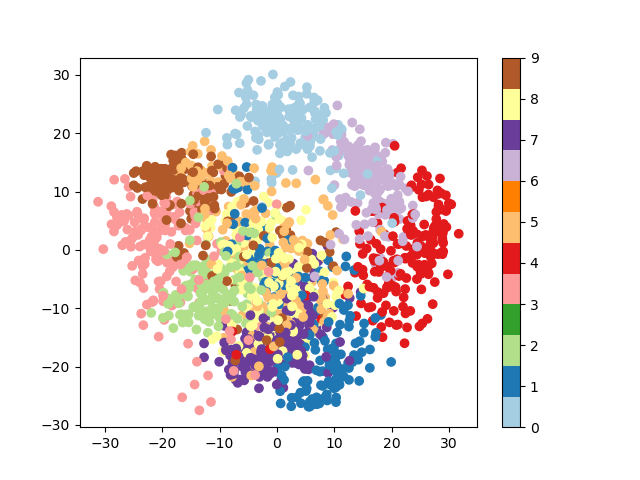
Question
Given these projections of the data, which numbers do you think a classifier might have trouble distinguishing?
three.six.three.iii. Gaussian Naive Bayes Classification¶
For most classification problems, it's nice to have a unproblematic, fast method to provide a quick baseline nomenclature. If the unproblematic and fast method is sufficient, then we don't have to waste product CPU cycles on more than circuitous models. If not, we can use the results of the simple method to give us clues about our data.
1 good method to keep in mind is Gaussian Naive Bayes ( sklearn.naive_bayes.GaussianNB ).
Tip
Gaussian Naive Bayes fits a Gaussian distribution to each training label independantly on each feature, and uses this to apace give a crude classification. It is generally not sufficiently accurate for real-globe information, merely can perform surprisingly well, for example on text data.
>>> from sklearn.naive_bayes import GaussianNB >>> from sklearn.model_selection import train_test_split >>> # separate the information into training and validation sets >>> X_train , X_test , y_train , y_test = train_test_split ( digits . information , digits . target ) >>> # train the model >>> clf = GaussianNB () >>> clf . fit ( X_train , y_train ) GaussianNB(priors=None) >>> # utilise the model to predict the labels of the test data >>> predicted = clf . predict ( X_test ) >>> expected = y_test >>> impress ( predicted ) [1 7 seven 7 8 two viii 0 iv viii vii 7 0 8 2 3 v 8 5 3 7 9 6 2 8 2 2 7 three 5...] >>> impress ( expected ) [1 0 iv 7 eight 2 ii 0 4 three seven 7 0 8 2 3 4 8 5 3 seven 9 half-dozen 3 8 2 2 9 3 v...] As higher up, we plot the digits with the predicted labels to get an thought of how well the nomenclature is working.

Question
Why did nosotros separate the data into training and validation sets?
iii.6.3.4. Quantitative Measurement of Functioning¶
We'd like to measure out the performance of our estimator without having to resort to plotting examples. A simple method might exist to simply compare the number of matches:
>>> matches = ( predicted == expected ) >>> print ( matches . sum ()) 367 >>> impress ( len ( matches )) 450 >>> matches . sum () / float ( len ( matches )) 0.81555555555555559 We come across that more than 80% of the 450 predictions lucifer the input. Simply there are other more sophisticated metrics that tin can be used to gauge the performance of a classifier: several are available in the sklearn.metrics submodule.
One of the most useful metrics is the classification_report , which combines several measures and prints a table with the results:
>>> from sklearn import metrics >>> impress ( metrics . classification_report ( expected , predicted )) precision recall f1-score support 0 1.00 0.91 0.95 46 ane 0.76 0.64 0.69 44 2 0.85 0.62 0.72 47 iii 0.98 0.82 0.89 49 4 0.89 0.86 0.88 37 v 0.97 0.93 0.95 41 6 ane.00 0.98 0.99 44 7 0.73 1.00 0.84 45 8 0.50 0.ninety 0.64 49 9 0.93 0.54 0.68 48 avg / total 0.86 0.82 0.82 450 Another enlightening metric for this sort of multi-characterization classification is a confusion matrix: it helps us visualize which labels are being interchanged in the classification errors:
>>> print ( metrics . confusion_matrix ( expected , predicted )) [[42 0 0 0 3 0 0 1 0 0] [ 0 28 0 0 0 0 0 1 13 2] [ 0 3 29 0 0 0 0 0 15 0] [ 0 0 ii 40 0 0 0 2 5 0] [ 0 0 1 0 32 ane 0 three 0 0] [ 0 0 0 0 0 38 0 2 1 0] [ 0 0 1 0 0 0 43 0 0 0] [ 0 0 0 0 0 0 0 45 0 0] [ 0 3 one 0 0 0 0 i 44 0] [ 0 three 0 ane 1 0 0 7 10 26]] We see here that in particular, the numbers 1, 2, 3, and 9 are often being labeled eight.
3.6.4. Supervised Learning: Regression of Housing Data¶
Here we'll do a short case of a regression problem: learning a continuous value from a set of features.
iii.6.4.one. A quick look at the data¶
We'll use the simple Boston house prices ready, bachelor in scikit-learn. This records measurements of 13 attributes of housing markets effectually Boston, as well equally the median cost. The question is: tin you predict the toll of a new market given its attributes?:
>>> from sklearn.datasets import load_boston >>> information = load_boston () >>> print ( information . data . shape ) (506, 13) >>> print ( data . target . shape ) (506,) Nosotros can see that there are merely over 500 data points.
The DESCR variable has a long description of the dataset:
>>> print ( data . DESCR ) Boston House Prices dataset =========================== Notes ------ Information Set up Characteristics: :Number of Instances: 506 :Number of Attributes: 13 numeric/categorical predictive :Median Value (aspect 14) is commonly the target :Aspect Information (in gild): - CRIM per capita criminal offence charge per unit by town - ZN proportion of residential land zoned for lots over 25,000 sq.ft. - INDUS proportion of not-retail business acres per town - CHAS Charles River dummy variable (= 1 if tract bounds river; 0 otherwise) - NOX nitric oxides concentration (parts per x million) - RM average number of rooms per dwelling - Historic period proportion of owner-occupied units built prior to 1940 - DIS weighted distances to five Boston employment centres - RAD index of accessibility to radial highways - Taxation full-value property-revenue enhancement charge per unit per $x,000 - PTRATIO student-teacher ratio by town - B m(Bk - 0.63)^2 where Bk is the proportion of blacks by boondocks - LSTAT % lower condition of the population - MEDV Median value of owner-occupied homes in $1000's ... It often helps to apace visualize pieces of the data using histograms, scatter plots, or other plot types. With matplotlib, allow us bear witness a histogram of the target values: the median price in each neighborhood:
>>> plt . hist ( data . target ) (array([... 
Let'due south accept a quick wait to come across if some features are more relevant than others for our problem:
>>> for index , feature_name in enumerate ( information . feature_names ): ... plt . effigy () ... plt . scatter ( data . data [:, index ], data . target ) <Effigy size... 












This is a manual version of a technique called feature choice.
Tip
Sometimes, in Automobile Learning it is useful to employ feature option to decide which features are the most useful for a particular problem. Automated methods exist which quantify this sort of practise of choosing the virtually informative features.
3.vi.4.2. Predicting Home Prices: a Simple Linear Regression¶
Now we'll use scikit-acquire to perform a uncomplicated linear regression on the housing data. At that place are many possibilities of regressors to use. A particularly uncomplicated one is LinearRegression : this is basically a wrapper effectually an ordinary least squares calculation.
>>> from sklearn.model_selection import train_test_split >>> X_train , X_test , y_train , y_test = train_test_split ( data . data , data . target ) >>> from sklearn.linear_model import LinearRegression >>> clf = LinearRegression () >>> clf . fit ( X_train , y_train ) LinearRegression(copy_X=True, fit_intercept=Truthful, n_jobs=one, normalize=False) >>> predicted = clf . predict ( X_test ) >>> expected = y_test >>> impress ( "RMS: %due south " % np . sqrt ( np . mean (( predicted - expected ) ** 2 ))) RMS: 5.0059... 
We can plot the mistake: expected equally a function of predicted:
>>> plt . scatter ( expected , predicted ) <matplotlib.collections.PathCollection object at ...> Tip
The prediction at least correlates with the true price, though at that place are clearly some biases. We could imagine evaluating the functioning of the regressor by, say, computing the RMS residuals betwixt the truthful and predicted toll. There are some subtleties in this, however, which we'll cover in a later on section.
Exercise: Gradient Boosting Tree Regression
At that place are many other types of regressors available in scikit-learn: nosotros'll endeavour a more powerful ane here.
Use the GradientBoostingRegressor class to fit the housing data.
hint You lot can copy and paste some of the higher up code, replacing LinearRegression with GradientBoostingRegressor :
from sklearn.ensemble import GradientBoostingRegressor # Instantiate the model, fit the results, and besprinkle in vs. out Solution The solution is plant in the code of this chapter
iii.6.v. Measuring prediction performance¶
3.half-dozen.5.one. A quick exam on the K-neighbors classifier¶
Here we'll continue to wait at the digits data, only we'll switch to the 1000-Neighbors classifier. The G-neighbors classifier is an example-based classifier. The K-neighbors classifier predicts the label of an unknown indicate based on the labels of the K nearest points in the parameter infinite.
>>> # Go the data >>> from sklearn.datasets import load_digits >>> digits = load_digits () >>> X = digits . data >>> y = digits . target >>> # Instantiate and train the classifier >>> from sklearn.neighbors import KNeighborsClassifier >>> clf = KNeighborsClassifier ( n_neighbors = ane ) >>> clf . fit ( 10 , y ) KNeighborsClassifier(...) >>> # Check the results using metrics >>> from sklearn import metrics >>> y_pred = clf . predict ( X ) >>> impress ( metrics . confusion_matrix ( y_pred , y )) [[178 0 0 0 0 0 0 0 0 0] [ 0 182 0 0 0 0 0 0 0 0] [ 0 0 177 0 0 0 0 0 0 0] [ 0 0 0 183 0 0 0 0 0 0] [ 0 0 0 0 181 0 0 0 0 0] [ 0 0 0 0 0 182 0 0 0 0] [ 0 0 0 0 0 0 181 0 0 0] [ 0 0 0 0 0 0 0 179 0 0] [ 0 0 0 0 0 0 0 0 174 0] [ 0 0 0 0 0 0 0 0 0 180]] Plain, nosotros've found a perfect classifier! But this is misleading for the reasons we saw before: the classifier essentially "memorizes" all the samples it has already seen. To really exam how well this algorithm does, we need to try some samples it hasn't still seen.
This trouble too occurs with regression models. In the following we fit an other instance-based model named "decision tree" to the Boston Housing toll dataset we introduced previously:
>>> from sklearn.datasets import load_boston >>> from sklearn.tree import DecisionTreeRegressor >>> information = load_boston () >>> clf = DecisionTreeRegressor () . fit ( information . data , data . target ) >>> predicted = clf . predict ( information . data ) >>> expected = data . target >>> plt . scatter ( expected , predicted ) <matplotlib.collections.PathCollection object at ...> >>> plt . plot ([ 0 , 50 ], [ 0 , fifty ], '--m' ) [<matplotlib.lines.Line2D object at ...] 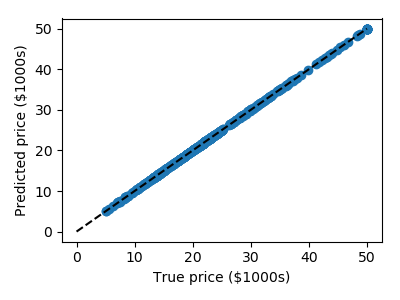
Hither once more the predictions are seemingly perfect as the model was able to perfectly memorize the preparation ready.
Warning
Performance on test fix
Operation on test prepare does not measure overfit (equally described above)
3.6.5.2. A correct approach: Using a validation set¶
Learning the parameters of a prediction function and testing it on the same information is a methodological mistake: a model that would just echo the labels of the samples that information technology has but seen would have a perfect score only would fail to predict annihilation useful on yet-unseen data.
To avoid over-fitting, we have to define two different sets:
- a training set X_train, y_train which is used for learning the parameters of a predictive model
- a testing prepare X_test, y_test which is used for evaluating the fitted predictive model
In scikit-learn such a random split can be quickly computed with the train_test_split() function:
>>> from sklearn import model_selection >>> X = digits . data >>> y = digits . target >>> X_train , X_test , y_train , y_test = model_selection . train_test_split ( X , y , ... test_size = 0.25 , random_state = 0 ) >>> print ( " %r , %r , %r " % ( X . shape , X_train . shape , X_test . shape )) (1797, 64), (1347, 64), (450, 64) At present we train on the training data, and test on the testing information:
>>> clf = KNeighborsClassifier ( n_neighbors = ane ) . fit ( X_train , y_train ) >>> y_pred = clf . predict ( X_test ) >>> impress ( metrics . confusion_matrix ( y_test , y_pred )) [[37 0 0 0 0 0 0 0 0 0] [ 0 43 0 0 0 0 0 0 0 0] [ 0 0 43 1 0 0 0 0 0 0] [ 0 0 0 45 0 0 0 0 0 0] [ 0 0 0 0 38 0 0 0 0 0] [ 0 0 0 0 0 47 0 0 0 one] [ 0 0 0 0 0 0 52 0 0 0] [ 0 0 0 0 0 0 0 48 0 0] [ 0 0 0 0 0 0 0 0 48 0] [ 0 0 0 1 0 one 0 0 0 45]] >>> print ( metrics . classification_report ( y_test , y_pred )) precision recall f1-score support 0 one.00 i.00 1.00 37 1 1.00 ane.00 i.00 43 ii 1.00 0.98 0.99 44 3 0.96 1.00 0.98 45 4 1.00 1.00 1.00 38 5 0.98 0.98 0.98 48 6 1.00 1.00 1.00 52 7 1.00 1.00 1.00 48 8 i.00 1.00 1.00 48 9 0.98 0.96 0.97 47 avg / total 0.99 0.99 0.99 450 The averaged f1-score is ofttimes used equally a convenient measure of the overall performance of an algorithm. It appears in the bottom row of the nomenclature report; it tin can besides be accessed directly:
>>> metrics . f1_score ( y_test , y_pred , average = "macro" ) 0.991367... The over-plumbing fixtures we saw previously can be quantified past calculating the f1-score on the training data itself:
>>> metrics . f1_score ( y_train , clf . predict ( X_train ), average = "macro" ) ane.0 Annotation
Regression metrics In the instance of regression models, we demand to use different metrics, such as explained variance.
three.6.5.3. Model Selection via Validation¶
Tip
We have practical Gaussian Naives, support vectors machines, and M-nearest neighbors classifiers to the digits dataset. Now that nosotros have these validation tools in identify, we can ask quantitatively which of the three estimators works best for this dataset.
-
With the default hyper-parameters for each estimator, which gives the all-time f1 score on the validation set? Call back that hyperparameters are the parameters set when you instantiate the classifier: for example, the
n_neighborsinclf = KNeighborsClassifier(n_neighbors=ane)>>> from sklearn.naive_bayes import GaussianNB >>> from sklearn.neighbors import KNeighborsClassifier >>> from sklearn.svm import LinearSVC >>> Ten = digits . data >>> y = digits . target >>> X_train , X_test , y_train , y_test = model_selection . train_test_split ( X , y , ... test_size = 0.25 , random_state = 0 ) >>> for Model in [ GaussianNB , KNeighborsClassifier , LinearSVC ]: ... clf = Model () . fit ( X_train , y_train ) ... y_pred = clf . predict ( X_test ) ... print ( ' %s : %s ' % ... ( Model . __name__ , metrics . f1_score ( y_test , y_pred , boilerplate = "macro" ))) GaussianNB: 0.8332741681... KNeighborsClassifier: 0.9804562804... LinearSVC: 0.93...
-
For each classifier, which value for the hyperparameters gives the best results for the digits data? For
LinearSVC, applyloss='l2'andloss='l1'. ForKNeighborsClassifierwe usen_neighborsbetween 1 and 10. Annotation thatGaussianNBdoes not have whatsoever adaptable hyperparameters.LinearSVC ( loss = 'l1' ): 0.930570687535 LinearSVC ( loss = 'l2' ): 0.933068826918 ------------------- KNeighbors ( n_neighbors = i ): 0.991367521884 KNeighbors ( n_neighbors = 2 ): 0.984844206884 KNeighbors ( n_neighbors = 3 ): 0.986775344954 KNeighbors ( n_neighbors = 4 ): 0.980371905382 KNeighbors ( n_neighbors = 5 ): 0.980456280495 KNeighbors ( n_neighbors = half dozen ): 0.975792419414 KNeighbors ( n_neighbors = seven ): 0.978064579214 KNeighbors ( n_neighbors = 8 ): 0.978064579214 KNeighbors ( n_neighbors = 9 ): 0.978064579214 KNeighbors ( n_neighbors = 10 ): 0.975555089773
Solution: code source
3.6.5.iv. Cross-validation¶
Cross-validation consists in repetively splitting the data in pairs of train and test sets, called 'folds'. Scikit-learn comes with a role to automatically compute score on all these folds. Here nosotros do KFold with g=5.
>>> clf = KNeighborsClassifier () >>> from sklearn.model_selection import cross_val_score >>> cross_val_score ( clf , 10 , y , cv = five ) array([0.9478022 , 0.9558011 , 0.96657382, 0.98039216, 0.96338028]) We can use dissimilar splitting strategies, such equally random splitting:
>>> from sklearn.model_selection import ShuffleSplit >>> cv = ShuffleSplit ( n_splits = v ) >>> cross_val_score ( clf , X , y , cv = cv ) array([...]) 3.6.5.5. Hyperparameter optimization with cross-validation¶
Consider regularized linear models, such as Ridge Regression, which uses l2 regularlization, and Lasso Regression, which uses l1 regularization. Choosing their regularization parameter is of import.
Let us set these parameters on the Diabetes dataset, a simple regression problem. The diabetes data consists of ten physiological variables (age, sexual activity, weight, claret pressure level) measure on 442 patients, and an indication of affliction progression later on one year:
>>> from sklearn.datasets import load_diabetes >>> data = load_diabetes () >>> 10 , y = data . information , data . target >>> print ( X . shape ) (442, 10) With the default hyper-parameters: we compute the cross-validation score:
>>> from sklearn.linear_model import Ridge , Lasso >>> for Model in [ Ridge , Lasso ]: ... model = Model () ... impress ( ' %s : %s ' % ( Model . __name__ , cross_val_score ( model , X , y ) . mean ())) Ridge: 0.409427438303 Lasso: 0.353800083299 Basic Hyperparameter Optimization¶
Nosotros compute the cross-validation score as a function of alpha, the force of the regularization for Lasso and Ridge . We choose 20 values of alpha between 0.0001 and 1:
>>> alphas = np . logspace ( - iii , - 1 , 30 ) >>> for Model in [ Lasso , Ridge ]: ... scores = [ cross_val_score ( Model ( alpha ), 10 , y , cv = iii ) . mean () ... for blastoff in alphas ] ... plt . plot ( alphas , scores , label = Model . __name__ ) [<matplotlib.lines.Line2D object at ... 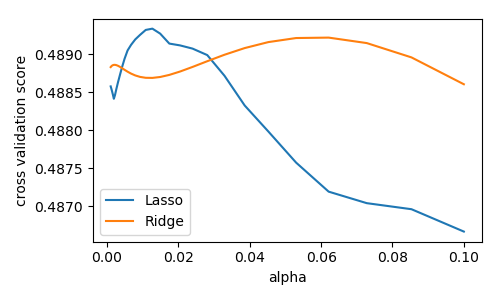
Question
Can we trust our results to exist actually useful?
Automatically Performing Grid Search¶
sklearn.grid_search.GridSearchCV is constructed with an estimator, too as a dictionary of parameter values to be searched. We can find the optimal parameters this way:
>>> from sklearn.grid_search import GridSearchCV >>> for Model in [ Ridge , Lasso ]: ... gscv = GridSearchCV ( Model (), dict ( alpha = alphas ), cv = 3 ) . fit ( X , y ) ... impress ( ' %southward : %s ' % ( Model . __name__ , gscv . best_params_ )) Ridge: {'blastoff': 0.062101694189156162} Lasso: {'blastoff': 0.01268961003167922} Born Hyperparameter Search¶
For some models inside scikit-acquire, cross-validation can be performed more efficiently on large datasets. In this case, a cross-validated version of the item model is included. The cantankerous-validated versions of Ridge and Lasso are RidgeCV and LassoCV , respectively. Parameter search on these estimators can be performed equally follows:
>>> from sklearn.linear_model import RidgeCV , LassoCV >>> for Model in [ RidgeCV , LassoCV ]: ... model = Model ( alphas = alphas , cv = 3 ) . fit ( X , y ) ... impress ( ' %south : %s ' % ( Model . __name__ , model . alpha_ )) RidgeCV: 0.0621016941892 LassoCV: 0.0126896100317 Nosotros run into that the results friction match those returned by GridSearchCV
Nested cross-validation¶
How practise we measure the operation of these estimators? We have used data to set the hyperparameters, so we demand to test on actually new data. We can practise this by running cross_val_score() on our CV objects. Here at that place are 2 cross-validation loops going on, this is called 'nested cross validation':
for Model in [ RidgeCV , LassoCV ]: scores = cross_val_score ( Model ( alphas = alphas , cv = 3 ), 10 , y , cv = 3 ) print ( Model . __name__ , np . mean ( scores )) Note
Note that these results do not match the all-time results of our curves above, and LassoCV seems to nether-perform RidgeCV . The reason is that setting the hyper-parameter is harder for Lasso, thus the estimation error on this hyper-parameter is larger.
3.vi.6. Unsupervised Learning: Dimensionality Reduction and Visualization¶
Unsupervised learning is applied on X without y: information without labels. A typical use instance is to detect hidden construction in the data.
3.6.six.one. Dimensionality Reduction: PCA¶
Dimensionality reduction derives a fix of new artificial features smaller than the original characteristic prepare. Hither we'll use Chief Component Analysis (PCA), a dimensionality reduction that strives to retain most of the variance of the original data. We'll use sklearn.decomposition.PCA on the iris dataset:
>>> Ten = iris . information >>> y = iris . target Tip
PCA computes linear combinations of the original features using a truncated Singular Value Decomposition of the matrix X, to projection the data onto a base of the top atypical vectors.
>>> from sklearn.decomposition import PCA >>> pca = PCA ( n_components = 2 , whiten = True ) >>> pca . fit ( Ten ) PCA(..., n_components=2, ...) Once fitted, PCA exposes the atypical vectors in the components_ attribute:
>>> pca . components_ array([[ 0.36158..., -0.08226..., 0.85657..., 0.35884...], [ 0.65653..., 0.72971..., -0.17576..., -0.07470...]]) Other attributes are available as well:
>>> pca . explained_variance_ratio_ array([0.92461..., 0.05301...]) Let usa projection the iris dataset along those first two dimensions::
>>> X_pca = pca . transform ( X ) >>> X_pca . shape (150, 2) PCA normalizes and whitens the data, which means that the information is at present centered on both components with unit of measurement variance:
>>> X_pca . hateful ( axis = 0 ) assortment([...eastward-15, ...e-15]) >>> X_pca . std ( centrality = 0 , ddof = 1 ) array([1., 1.]) Furthermore, the samples components do no longer carry whatever linear correlation:
>>> np . corrcoef ( X_pca . T ) array([[1.00000000e+00, 0.0], [0.0, 1.00000000e+00]]) With a number of retained components 2 or 3, PCA is useful to visualize the dataset:
>>> target_ids = range ( len ( iris . target_names )) >>> for i , c , label in zip ( target_ids , 'rgbcmykw' , iris . target_names ): ... plt . scatter ( X_pca [ y == i , 0 ], X_pca [ y == i , i ], ... c = c , label = label ) <matplotlib.collections.PathCollection ... 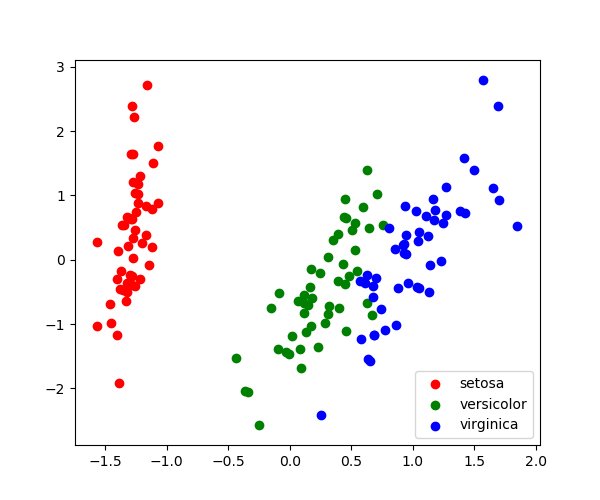
Tip
Note that this projection was determined without any information about the labels (represented by the colors): this is the sense in which the learning is unsupervised. Nevertheless, nosotros run across that the projection gives the states insight into the distribution of the different flowers in parameter infinite: notably, iris setosa is much more than distinct than the other two species.
3.six.6.two. Visualization with a non-linear embedding: tSNE¶
For visualization, more complex embeddings can be useful (for statistical analysis, they are harder to control). sklearn.manifold.TSNE is such a powerful manifold learning method. Nosotros apply it to the digits dataset, as the digits are vectors of dimension 8*eight = 64. Embedding them in 2d enables visualization:
>>> # Accept the starting time 500 information points: it'south difficult to see 1500 points >>> X = digits . data [: 500 ] >>> y = digits . target [: 500 ] >>> # Fit and transform with a TSNE >>> from sklearn.manifold import TSNE >>> tsne = TSNE ( n_components = 2 , random_state = 0 ) >>> X_2d = tsne . fit_transform ( X ) >>> # Visualize the data >>> plt . besprinkle ( X_2d [:, 0 ], X_2d [:, i ], c = y ) <matplotlib.collections.PathCollection object at ...> 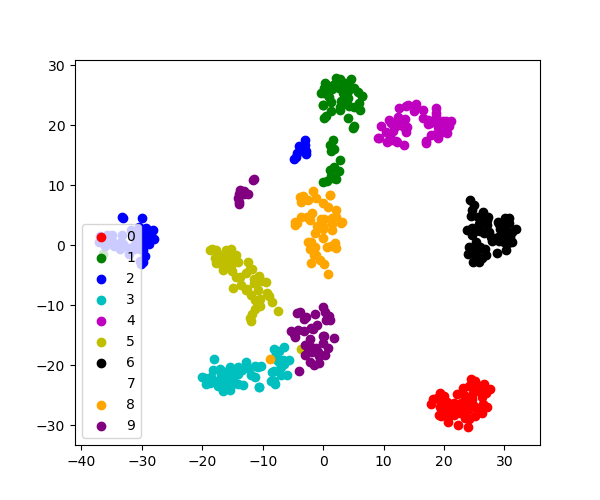
fit_transform
As TSNE cannot be applied to new information, we demand to utilise its fit_transform method.
sklearn.manifold.TSNE separates quite well the dissimilar classes of digits eventhough it had no access to the class information.
Practise: Other dimension reduction of digits
sklearn.manifold has many other not-linear embeddings. Try them out on the digits dataset. Could you judge their quality without knowing the labels y ?
>>> from sklearn.datasets import load_digits >>> digits = load_digits () >>> # ... 3.6.8. The eigenfaces instance: chaining PCA and SVMs¶
The goal of this example is to bear witness how an unsupervised method and a supervised one can be chained for better prediction. It starts with a didactic but lengthy way of doing things, and finishes with the idiomatic arroyo to pipelining in scikit-learn.
Here we'll take a expect at a unproblematic facial recognition instance. Ideally, we would use a dataset consisting of a subset of the Labeled Faces in the Wild data that is available with sklearn.datasets.fetch_lfw_people() . Even so, this is a relatively large download (~200MB) so nosotros volition exercise the tutorial on a simpler, less rich dataset. Feel free to explore the LFW dataset.
from sklearn import datasets faces = datasets . fetch_olivetti_faces () faces . data . shape Let'southward visualize these faces to run into what we're working with
from matplotlib import pyplot as plt fig = plt . figure ( figsize = ( viii , six )) # plot several images for i in range ( 15 ): ax = fig . add_subplot ( iii , 5 , i + 1 , xticks = [], yticks = []) ax . imshow ( faces . images [ i ], cmap = plt . cm . os )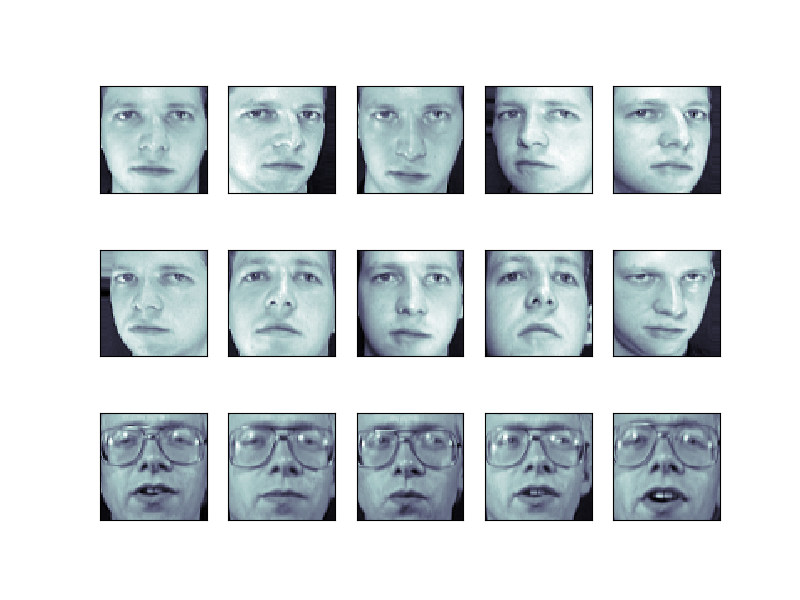 
Tip
Note is that these faces take already been localized and scaled to a common size. This is an important preprocessing piece for facial recognition, and is a procedure that can crave a large drove of grooming information. This can be done in scikit-learn, just the challenge is gathering a sufficient amount of preparation data for the algorithm to work. Fortunately, this slice is mutual enough that it has been done. I good resources is OpenCV, the Open Computer Vision Library.
Nosotros'll perform a Support Vector nomenclature of the images. We'll exercise a typical train-examination separate on the images:
from sklearn.model_selection import train_test_split X_train , X_test , y_train , y_test = train_test_split ( faces . information , faces . target , random_state = 0 ) print ( X_train . shape , X_test . shape ) Out:
3.6.eight.1. Preprocessing: Primary Component Analysis¶
1850 dimensions is a lot for SVM. Nosotros tin use PCA to reduce these 1850 features to a manageable size, while maintaining most of the information in the dataset.
from sklearn import decomposition pca = decomposition . PCA ( n_components = 150 , whiten = True ) pca . fit ( X_train ) One interesting part of PCA is that information technology computes the "mean" face, which can be interesting to examine:
plt . imshow ( pca . mean_ . reshape ( faces . images [ 0 ] . shape ), cmap = plt . cm . os ) 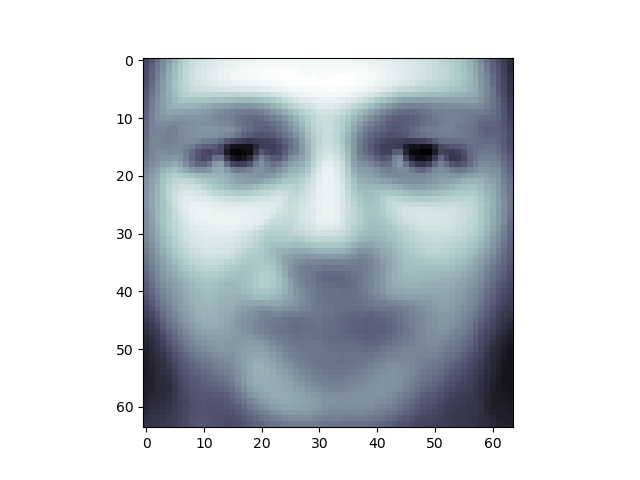
The principal components measure deviations about this mean along orthogonal axes.
print ( pca . components_ . shape ) Out:
It is also interesting to visualize these principal components:
fig = plt . figure ( figsize = ( xvi , 6 )) for i in range ( xxx ): ax = fig . add_subplot ( 3 , 10 , i + 1 , xticks = [], yticks = []) ax . imshow ( pca . components_ [ i ] . reshape ( faces . images [ 0 ] . shape ), cmap = plt . cm . bone )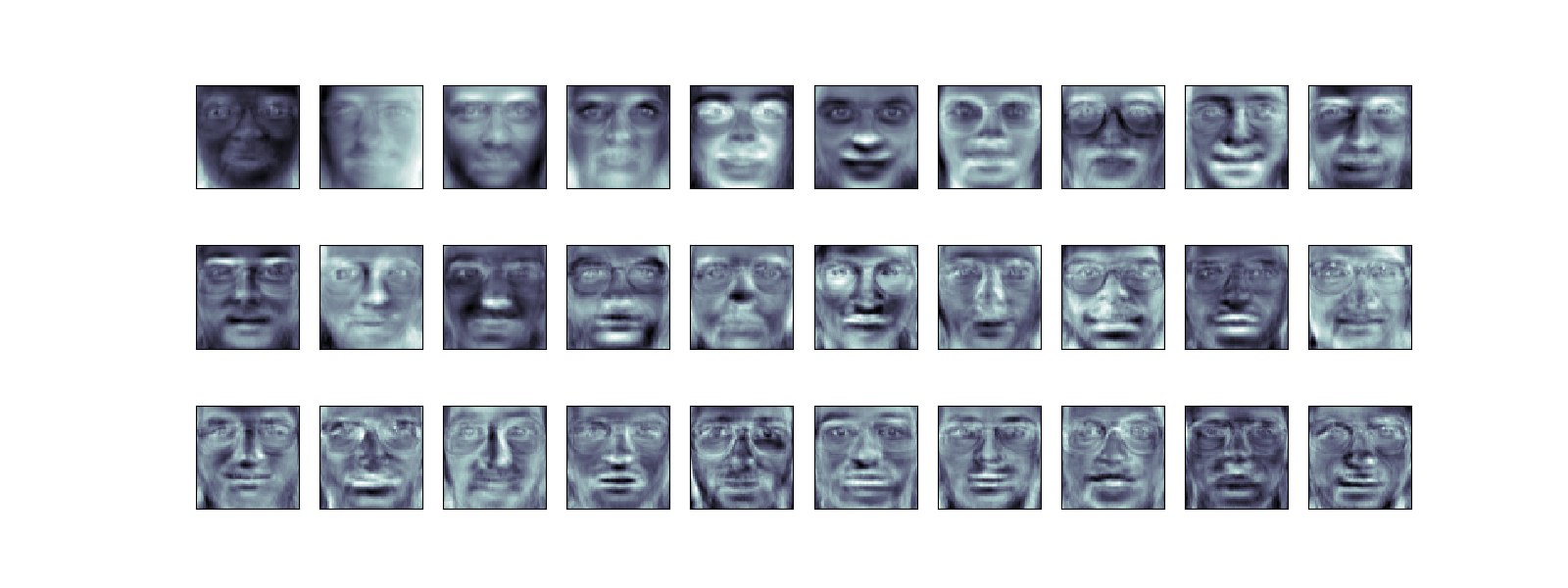 
The components ("eigenfaces") are ordered past their importance from tiptop-left to bottom-right. Nosotros see that the first few components seem to primarily accept care of lighting conditions; the remaining components pull out certain identifying features: the nose, eyes, eyebrows, etc.
With this project computed, we can now project our original training and test information onto the PCA basis:
X_train_pca = pca . transform ( X_train ) X_test_pca = pca . transform ( X_test ) impress ( X_train_pca . shape ) Out:
Out:
These projected components correspond to factors in a linear combination of component images such that the combination approaches the original face.
three.6.8.two. Doing the Learning: Support Vector Machines¶
Now we'll perform support-vector-motorcar classification on this reduced dataset:
from sklearn import svm clf = svm . SVC ( C = 5. , gamma = 0.001 ) clf . fit ( X_train_pca , y_train ) Finally, we can evaluate how well this classification did. First, we might plot a few of the examination-cases with the labels learned from the preparation prepare:
import numpy every bit np fig = plt . figure ( figsize = ( 8 , 6 )) for i in range ( 15 ): ax = fig . add_subplot ( 3 , 5 , i + 1 , xticks = [], yticks = []) ax . imshow ( X_test [ i ] . reshape ( faces . images [ 0 ] . shape ), cmap = plt . cm . os ) y_pred = clf . predict ( X_test_pca [ i , np . newaxis ])[ 0 ] color = ( 'black' if y_pred == y_test [ i ] else 'cerise' ) ax . set_title ( y_pred , fontsize = 'pocket-sized' , color = color )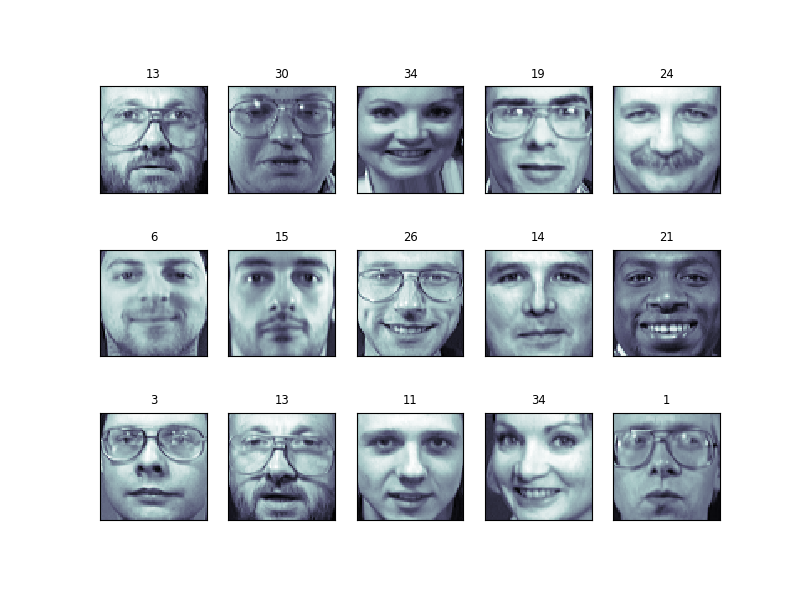 
The classifier is correct on an impressive number of images given the simplicity of its learning model! Using a linear classifier on 150 features derived from the pixel-level data, the algorithm correctly identifies a large number of the people in the images.
Again, we can quantify this effectiveness using one of several measures from sklearn.metrics . Beginning we can do the nomenclature report, which shows the precision, recall and other measures of the "goodness" of the classification:
from sklearn import metrics y_pred = clf . predict ( X_test_pca ) print ( metrics . classification_report ( y_test , y_pred )) Out:
precision think f1-score back up 0 ane.00 0.67 0.80 vi 1 ane.00 1.00 one.00 4 ii 0.fifty 1.00 0.67 2 three 1.00 ane.00 1.00 1 4 0.50 ane.00 0.67 1 5 1.00 1.00 i.00 v 6 1.00 1.00 one.00 4 7 1.00 0.67 0.eighty three 9 1.00 1.00 ane.00 1 x ane.00 1.00 one.00 4 eleven ane.00 1.00 1.00 ane 12 1.00 ane.00 one.00 ii thirteen i.00 1.00 1.00 3 xiv 1.00 one.00 1.00 v 15 0.75 i.00 0.86 3 17 1.00 1.00 1.00 half dozen xix one.00 1.00 1.00 4 20 1.00 1.00 ane.00 ane 21 1.00 1.00 1.00 1 22 1.00 ane.00 1.00 2 23 one.00 1.00 1.00 1 24 1.00 1.00 1.00 2 25 1.00 0.50 0.67 two 26 1.00 0.75 0.86 4 27 1.00 one.00 1.00 1 28 0.67 1.00 0.fourscore two 29 1.00 ane.00 ane.00 three 30 1.00 1.00 1.00 4 31 1.00 ane.00 1.00 3 32 i.00 1.00 1.00 iii 33 1.00 one.00 1.00 two 34 i.00 i.00 i.00 3 35 ane.00 1.00 1.00 i 36 1.00 i.00 1.00 3 37 1.00 1.00 1.00 3 38 1.00 1.00 1.00 1 39 i.00 i.00 1.00 3 avg / total 0.97 0.95 0.95 100 Another interesting metric is the confusion matrix, which indicates how oftentimes any two items are mixed-upwardly. The confusion matrix of a perfect classifier would only have nonzero entries on the diagonal, with zeros on the off-diagonal:
print ( metrics . confusion_matrix ( y_test , y_pred )) Out:
[[4 0 0 ... 0 0 0] [0 4 0 ... 0 0 0] [0 0 2 ... 0 0 0] ... [0 0 0 ... iii 0 0] [0 0 0 ... 0 1 0] [0 0 0 ... 0 0 3]] 3.6.eight.three. Pipelining¶
Higher up nosotros used PCA as a pre-processing step before applying our back up vector machine classifier. Plugging the output of one estimator directly into the input of a second figurer is a normally used design; for this reason scikit-learn provides a Pipeline object which automates this process. The above problem can be re-expressed equally a pipeline as follows:
from sklearn.pipeline import Pipeline clf = Pipeline ([( 'pca' , decomposition . PCA ( n_components = 150 , whiten = Truthful )), ( 'svm' , svm . LinearSVC ( C = 1.0 ))]) clf . fit ( X_train , y_train ) y_pred = clf . predict ( X_test ) print ( metrics . confusion_matrix ( y_pred , y_test )) iii.6.9. Parameter selection, Validation, and Testing¶
3.6.9.i. Hyperparameters, Over-fitting, and Under-fitting¶
The issues associated with validation and cross-validation are some of the most important aspects of the practice of machine learning. Selecting the optimal model for your information is vital, and is a slice of the trouble that is not frequently appreciated past auto learning practitioners.
The key question is: If our estimator is underperforming, how should we motility forward?
- Utilise simpler or more complicated model?
- Add together more features to each observed data point?
- Add more preparation samples?
The answer is oftentimes counter-intuitive. In particular, Sometimes using a more than complicated model will give worse results. Too, Sometimes calculation grooming data will non improve your results. The ability to decide what steps volition improve your model is what separates the successful motorcar learning practitioners from the unsuccessful.
Bias-variance trade-off: illustration on a simple regression problem¶
Let u.s.a. kickoff with a simple 1D regression problem. This will help u.s. to easily visualize the information and the model, and the results generalize easily to higher-dimensional datasets. We'll explore a simple linear regression problem, with sklearn.linear_model .
X = np . c_ [ . 5 , 1 ] . T y = [ . 5 , one ] X_test = np . c_ [ 0 , ii ] . T Without racket, every bit linear regression fits the data perfectly
from sklearn import linear_model regr = linear_model . LinearRegression () regr . fit ( 10 , y ) plt . plot ( 10 , y , 'o' ) plt . plot ( X_test , regr . predict ( X_test )) 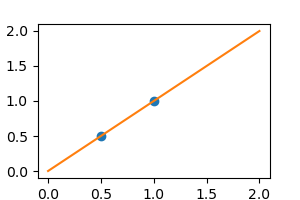
In real life situation, we accept noise (e.g. measurement noise) in our data:
np . random . seed ( 0 ) for _ in range ( half-dozen ): noisy_X = X + np . random . normal ( loc = 0 , calibration =. 1 , size = 10 . shape ) plt . plot ( noisy_X , y , 'o' ) regr . fit ( noisy_X , y ) plt . plot ( X_test , regr . predict ( X_test )) 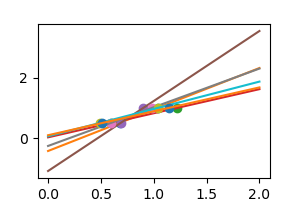
Every bit nosotros tin can see, our linear model captures and amplifies the racket in the data. It displays a lot of variance.
Nosotros tin employ some other linear estimator that uses regularization, the Ridge estimator. This figurer regularizes the coefficients by shrinking them to zero, under the supposition that very high correlations are ofttimes spurious. The blastoff parameter controls the corporeality of shrinkage used.
regr = linear_model . Ridge ( alpha =. 1 ) np . random . seed ( 0 ) for _ in range ( half-dozen ): noisy_X = X + np . random . normal ( loc = 0 , scale =. 1 , size = X . shape ) plt . plot ( noisy_X , y , 'o' ) regr . fit ( noisy_X , y ) plt . plot ( X_test , regr . predict ( X_test )) plt . bear witness () 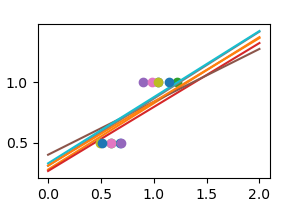
As we tin can see, the computer displays much less variance. Notwithstanding it systematically under-estimates the coefficient. Information technology displays a biased beliefs.
This is a typical example of bias/variance tradeof: non-regularized estimator are not biased, but they can display a lot of variance. Highly-regularized models have little variance, but high bias. This bias is non necessarily a bad matter: what matters is choosing the tradeoff between bias and variance that leads to the best prediction functioning. For a specific dataset there is a sweet spot respective to the highest complexity that the information can support, depending on the corporeality of noise and of observations available.
three.6.9.2. Visualizing the Bias/Variance Tradeoff¶
Tip
Given a item dataset and a model (east.g. a polynomial), we'd like to sympathize whether bias (underfit) or variance limits prediction, and how to melody the hyperparameter (hither d , the caste of the polynomial) to give the best fit.
On a given information, let us fit a uncomplicated polynomial regression model with varying degrees:
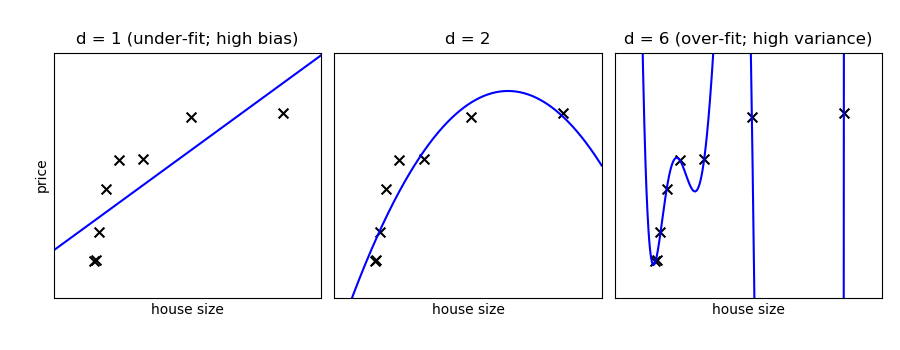
Tip
In the above figure, we see fits for three different values of d . For d = 1 , the data is under-fit. This means that the model is also simplistic: no straight line will ever be a proficient fit to this data. In this instance, nosotros say that the model suffers from high bias. The model itself is biased, and this volition be reflected in the fact that the data is poorly fit. At the other extreme, for d = vi the information is over-fit. This means that the model has too many free parameters (6 in this instance) which tin can be adjusted to perfectly fit the training data. If we add together a new point to this plot, though, chances are information technology will be very far from the curve representing the degree-6 fit. In this case, we say that the model suffers from high variance. The reason for the term "loftier variance" is that if any of the input points are varied slightly, it could result in a very unlike model.
In the middle, for d = ii , we accept found a good mid-point. It fits the data fairly well, and does not suffer from the bias and variance problems seen in the figures on either side. What we would like is a fashion to quantitatively identify bias and variance, and optimize the metaparameters (in this case, the polynomial degree d) in order to determine the best algorithm.
Polynomial regression with scikit-learn
A polynomial regression is built by pipelining PolynomialFeatures and a LinearRegression :
>>> from sklearn.pipeline import make_pipeline >>> from sklearn.preprocessing import PolynomialFeatures >>> from sklearn.linear_model import LinearRegression >>> model = make_pipeline ( PolynomialFeatures ( degree = ii ), LinearRegression ()) Validation Curves¶
Let us create a dataset like in the instance in a higher place:
>>> def generating_func ( x , err = 0.5 ): ... render np . random . normal ( 10 - 1. / ( x + 0.one ), err ) >>> # randomly sample more than data >>> np . random . seed ( 1 ) >>> x = np . random . random ( size = 200 ) >>> y = generating_func ( x , err = 1. ) 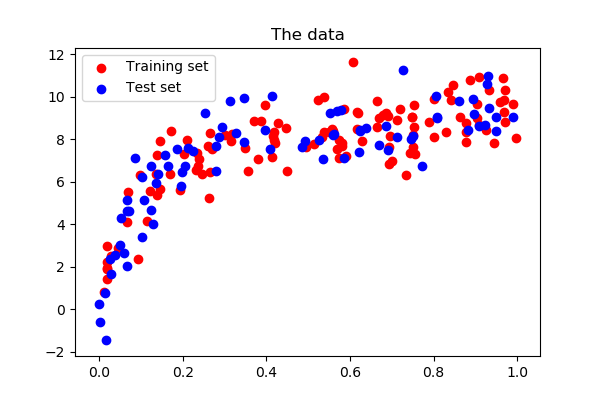
Central to quantify bias and variance of a model is to employ it on test data, sampled from the same distribution as the train, merely that will capture independent noise:
>>> xtrain , xtest , ytrain , ytest = train_test_split ( x , y , test_size = 0.4 ) Validation curve A validation bend consists in varying a model parameter that controls its complexity (here the caste of the polynomial) and measures both error of the model on training data, and on test information (eg with cantankerous-validation). The model parameter is then adapted so that the test mistake is minimized:
We use sklearn.model_selection.validation_curve() to compute railroad train and test fault, and plot it:
>>> from sklearn.model_selection import validation_curve >>> degrees = np . arange ( 1 , 21 ) >>> model = make_pipeline ( PolynomialFeatures (), LinearRegression ()) >>> # Vary the "degrees" on the pipeline pace "polynomialfeatures" >>> train_scores , validation_scores = validation_curve ( ... model , ten [:, np . newaxis ], y , ... param_name = 'polynomialfeatures__degree' , ... param_range = degrees ) >>> # Plot the mean railroad train score and validation score across folds >>> plt . plot ( degrees , validation_scores . mean ( centrality = 1 ), label = 'cross-validation' ) [<matplotlib.lines.Line2D object at ...>] >>> plt . plot ( degrees , train_scores . mean ( axis = 1 ), characterization = 'training' ) [<matplotlib.lines.Line2D object at ...>] >>> plt . legend ( loc = 'best' ) <matplotlib.fable.Legend object at ...> 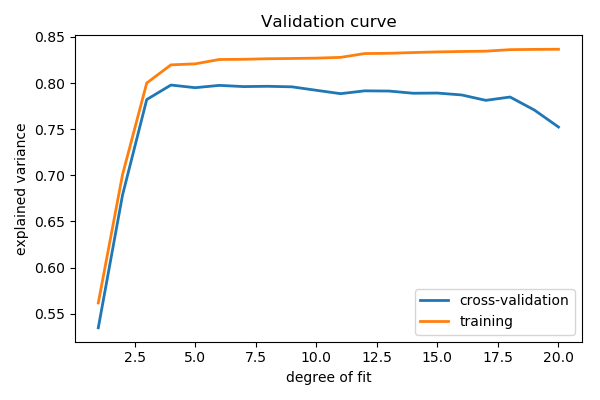
This figure shows why validation is important. On the left side of the plot, we have very low-caste polynomial, which under-fit the information. This leads to a low explained variance for both the training set and the validation set up. On the far right side of the plot, we have a very high degree polynomial, which over-fits the data. This tin be seen in the fact that the training explained variance is very loftier, while on the validation set up, it is low. Choosing d around iv or 5 gets us the best tradeoff.
Tip
The astute reader will realize that something is amiss here: in the above plot, d = 4 gives the all-time results. But in the previous plot, we establish that d = vi vastly over-fits the data. What's going on hither? The difference is the number of training points used. In the previous instance, in that location were simply eight training points. In this example, we accept 100. As a full general rule of thumb, the more training points used, the more complicated model tin exist used. But how can you determine for a given model whether more grooming points will be helpful? A useful diagnostic for this are learning curves.
Learning Curves¶
A learning curve shows the grooming and validation score equally a function of the number of preparation points. Note that when we railroad train on a subset of the training data, the grooming score is computed using this subset, not the full training set. This curve gives a quantitative view into how beneficial it will be to add training samples.
Questions:
- As the number of training samples are increased, what do yous expect to see for the training score? For the validation score?
- Would you expect the training score to exist higher or lower than the validation score? Would you ever expect this to change?
scikit-learn provides sklearn.model_selection.learning_curve() :
>>> from sklearn.model_selection import learning_curve >>> train_sizes , train_scores , validation_scores = learning_curve ( ... model , x [:, np . newaxis ], y , train_sizes = np . logspace ( - ane , 0 , 20 )) >>> # Plot the mean train score and validation score across folds >>> plt . plot ( train_sizes , validation_scores . mean ( axis = 1 ), characterization = 'cross-validation' ) [<matplotlib.lines.Line2D object at ...>] >>> plt . plot ( train_sizes , train_scores . hateful ( axis = 1 ), label = 'preparation' ) [<matplotlib.lines.Line2D object at ...>] 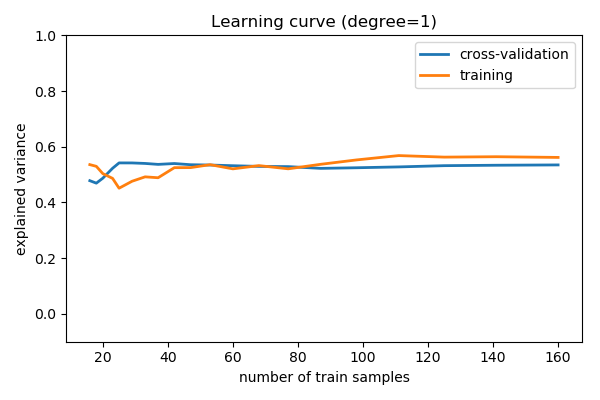
For a degree=i model
Note that the validation score generally increases with a growing training set, while the training score generally decreases with a growing training set. As the training size increases, they volition converge to a unmarried value.
From the in a higher place word, we know that d = one is a high-bias estimator which under-fits the information. This is indicated by the fact that both the preparation and validation scores are low. When confronted with this type of learning curve, we can expect that calculation more training data will not aid: both lines converge to a relatively low score.
When the learning curves accept converged to a depression score, we accept a high bias model.
A loftier-bias model can exist improved by:
- Using a more sophisticated model (i.e. in this case, increase
d) - Gather more features for each sample.
- Decrease regularization in a regularized model.
Increasing the number of samples, even so, does not improve a high-bias model.
Now let's look at a high-variance (i.eastward. over-fit) model:
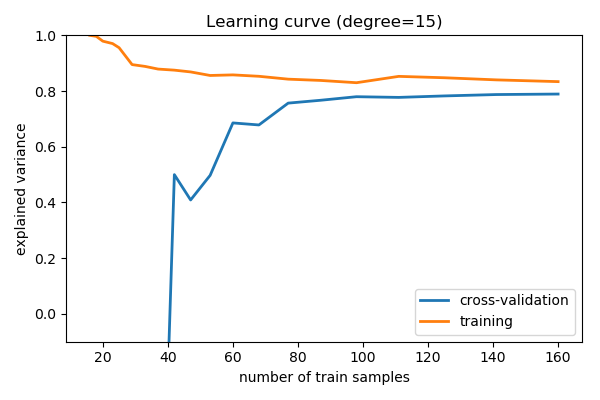
For a caste=xv model
Here we testify the learning bend for d = xv . From the above word, we know that d = 15 is a high-variance figurer which over-fits the data. This is indicated by the fact that the preparation score is much college than the validation score. As we add more samples to this grooming prepare, the training score volition continue to decrease, while the cross-validation fault will continue to increase, until they encounter in the middle.
Learning curves that accept not yet converged with the full preparation set indicate a loftier-variance, over-fit model.
A loftier-variance model can be improved past:
- Gathering more training samples.
- Using a less-sophisticated model (i.e. in this case, make
dsmaller) - Increasing regularization.
In item, gathering more than features for each sample will non help the results.
3.6.9.3. Summary on model pick¶
We've seen above that an under-performing algorithm tin exist due to ii possible situations: high bias (under-plumbing fixtures) and high variance (over-fitting). In guild to evaluate our algorithm, we gear up aside a portion of our training data for cross-validation. Using the technique of learning curves, we can train on progressively larger subsets of the data, evaluating the training error and cross-validation error to determine whether our algorithm has loftier variance or high bias. But what do we do with this information?
Loftier Bias¶
If a model shows loftier bias, the following actions might help:
- Add together more features. In our case of predicting home prices, it may be helpful to make use of information such equally the neighborhood the firm is in, the year the business firm was built, the size of the lot, etc. Adding these features to the training and test sets can improve a high-bias estimator
- Use a more than sophisticated model. Adding complication to the model can assist improve on bias. For a polynomial fit, this can be achieved by increasing the caste d. Each learning technique has its own methods of adding complexity.
- Employ fewer samples. Though this will not ameliorate the nomenclature, a loftier-bias algorithm can attain nearly the same error with a smaller training sample. For algorithms which are computationally expensive, reducing the preparation sample size can atomic number 82 to very big improvements in speed.
- Decrease regularization. Regularization is a technique used to impose simplicity in some machine learning models, by calculation a penalisation term that depends on the characteristics of the parameters. If a model has high bias, decreasing the consequence of regularization tin atomic number 82 to ameliorate results.
High Variance¶
If a model shows loftier variance, the post-obit deportment might aid:
- Employ fewer features. Using a feature choice technique may be useful, and decrease the over-fitting of the estimator.
- Use a simpler model. Model complexity and over-plumbing equipment go hand-in-paw.
- Apply more than grooming samples. Adding training samples can reduce the issue of over-plumbing fixtures, and atomic number 82 to improvements in a high variance computer.
- Increase Regularization. Regularization is designed to prevent over-fitting. In a loftier-variance model, increasing regularization tin can pb to better results.
These choices become very important in real-world situations. For case, due to limited telescope time, astronomers must seek a balance between observing a large number of objects, and observing a large number of features for each object. Determining which is more than important for a particular learning job tin can inform the observing strategy that the astronomer employs.
3.6.9.iv. A last give-and-take of caution: divide validation and exam set¶
Using validation schemes to decide hyper-parameters means that we are fitting the hyper-parameters to the particular validation set. In the same fashion that parameters can be over-fit to the training gear up, hyperparameters can exist over-fit to the validation set. Considering of this, the validation fault tends to under-predict the classification error of new data.
For this reason, it is recommended to split up the data into 3 sets:
- The training ready, used to train the model (normally ~60% of the data)
- The validation fix, used to validate the model (usually ~xx% of the data)
- The test set up, used to evaluate the expected mistake of the validated model (usually ~xx% of the information)
Many automobile learning practitioners do non split exam set and validation fix. But if your goal is to estimate the mistake of a model on unknown information, using an independent exam set is vital.
Source: https://scipy-lectures.org/packages/scikit-learn/index.html
0 Response to "Find the Orthogonal Trajectories of the Family of Curves. (Use C for Any Needed Constant.) Y2 = 5kx3"
Post a Comment